 to think about and work out the following:
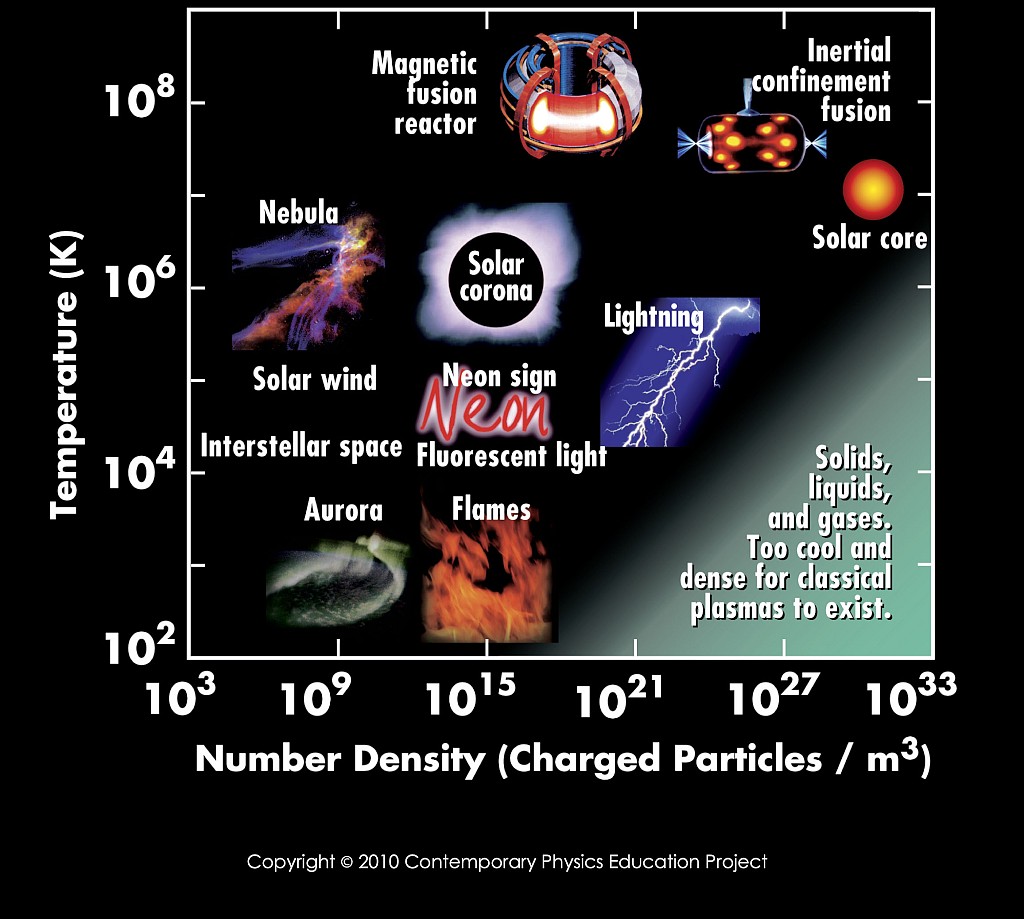
to think about and work out the following:As discussed in the Lecture, it is important to know the length scale on which the basic constituents of a medium interact. Start with the figure
 to think about and work out the following:
to think about and work out the following:
To make things as easy as possible for the numerical experiments, we prepared a mechanism so that you can run the numerical experiments on your laptop with minimal configuration efforts. We therefore prepared a docker image for you which you can run on your laptop to have the according environment.
So here some first steps to prepare:
vi
.tar file of the container you need for your system an place it in an extra work folder.
You can perform all tasks within your docker session and access the results from outside.
To do each tutorial, you need to follow some basic steps:
xterm if you have Linux/Mac or PowerShell if you have Windows)docker load -i opengadget3-container-XXX.tar, replace the XXX with what you need on your system)docker run -it -v .:/mnt --name opengadget3_container opengadget3:XXX)/mnt then it is visible from outside the container. (e.g. first mkdir /mnt/T01, then cd /mnt/T01) cp -r ~/OpenGadget3 . within the directory for your tutorial.cd OpenGadget3 wget https://www.usm.uni-muenchen.de/~dolag/Hydro/Hydro/Config.sh cd ..wget https://www.usm.uni-muenchen.de/~dolag/Hydro/Hydro/box.param wget https://www.usm.uni-muenchen.de/~dolag/Hydro/Hydro/box.iccd OpenGadget3 vi Config.sh (Note that this time you do not need to change anything) source Build/Container_build.sh (Note that this has to be done only once for each session)make -jcd ..vi box.param (Note that again, for this tutorial there is nothing to change)mpiexec -np 2 OpenGadget3/OpenGadget3 box.param wget https://www.usm.uni-muenchen.de/~dolag/Hydro/Hydro/Python/show_single.py vi show_single.py (Note that for the first step, there is nothing to change)python3 show_single.py
Congratulations, you have for the first time runned a full cycle of setting up, unnimng and analyzing a simulation. Now get a little bit more familiar with some details of the steps.
Have a more detailed look at the plotting script show_single.py which is kept quite simple.
You can identify the line where the output file snap_000 is selected, and where the positions pos are read from the file. From the scatter plot command you can see, that
pos is an array for all particles (first index) and the second index with encodes the x,y and z components. From the reading you can see that the label for the positions is "POS ".
Now you can try to read other vales of the particles from the file. You can try to use the velocity ("VEL ") which has three components like the positions before, or some scalar values like
the density ("RHO "), the internal energy ("U ") or the smoothing length of each particle ("HSML"). Names are always filled with SPACE so that they have 4
characters. Now you can experiment with different kind of scatter plots.
wget https://www.usm.uni-muenchen.de/~dolag/Hydro/Hydro/Python/show.py
python3 show.py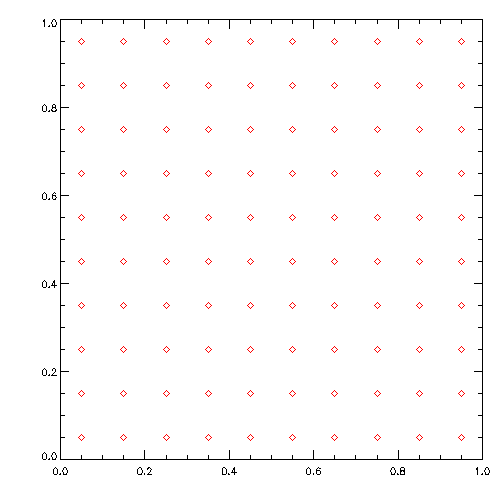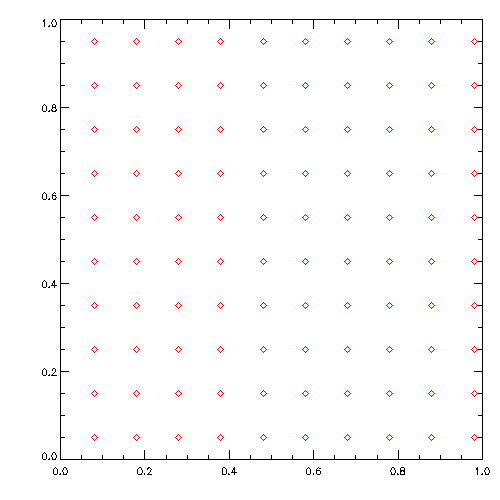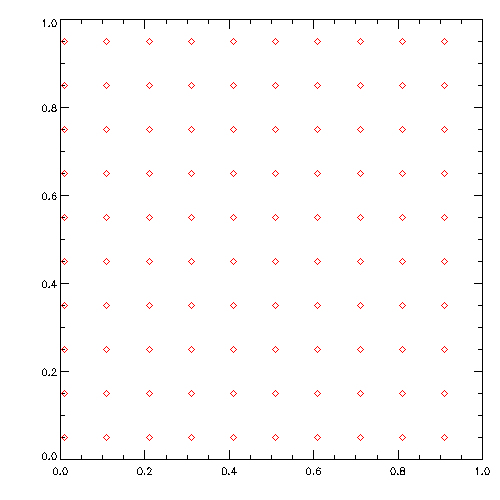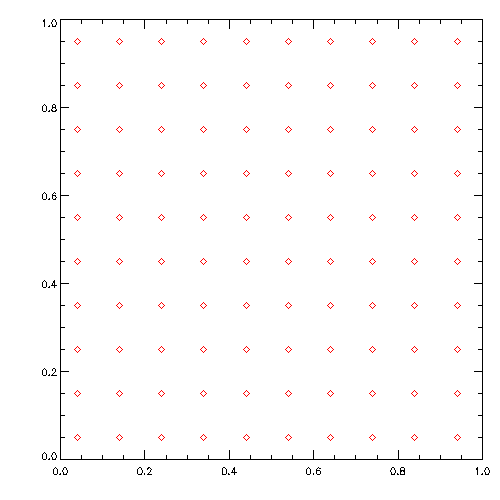
Initial conditions a basicly exactly the same files than the simulation output. They just contain all intrinsic properties (including the spacial distribution) of all SPH particles in the simulation. Starting from an empty template file ic_header we can just define arrays for the basic properties of the particles and assign them to define the geometry and setup of the experiment we want to do. As soon as we have done this, we can create a new box.ic file and do simulation based on this initial condition file.
So here an example, how to create the initial conditions for the problem we did before:
wget https://www.usm.uni-muenchen.de/~dolag/Hydro/Hydro/Python/setup_grid.py (This is an example script)
wget https://www.usm.uni-muenchen.de/~dolag/Hydro/Hydro/Python/ic_header (This has to be done only once to have the template file)
vi setup_grid.py (Let's have a look at the code, but nothing to change at the moment)
python3 setup_grid.pyvi setup_grid.py (Now change the direction of motion in the initial conditions)
python3 setup_grid.py>Goal of this tutorial is that you learn how to use commands in a Unix shell,
how to compile and execute a program,
how to create initial condition,
how to start a simulation, and
how to plot the results.